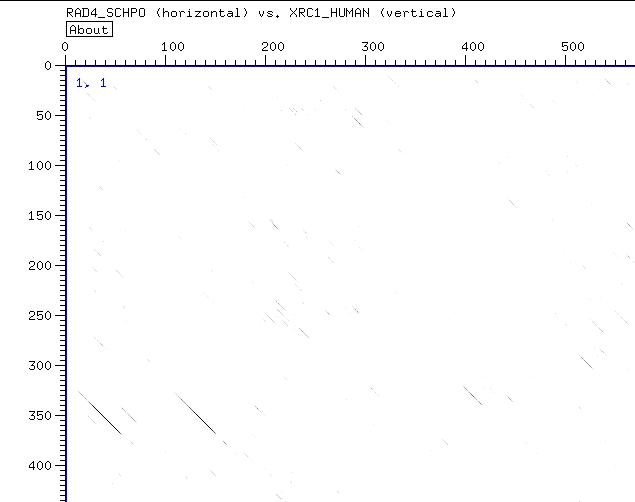
Fetching the sequences
Extracting Homologous Segments
Making an Alignment
- Cut and paste the sequence in the multiple alignment server of your choice
- Here is an alignment of the two segments: [two]
- Here is an alignment of the three segments: [three]
- The difference between the two alignments are obvious:[comparison]
Testing the validity of the model
- Add extra sequences to check if they affect the original alignment
- Extra sequences will be found using blast
- Sequences must be as diverse as possible, with key residues well conserved
- Here is an example of a correct alignment [ref_aln]
- Observe that also the sequences are distantly related they remain conserved on key positions
CONCLUSION
Although the similarity between hmga_chite.fasta and hmgl_trybr.fasta is clearly
within the twilight zone, the multiple alignment shows that they share key positions of the hmg
domain. As a consequence it is quite likely that these parts of the proteins are indeed homologous.
|